Spatial transcriptomics
Last updated: March 20, 2026
⚠ Spatial transcriptomics is currently available as an early Beta feature. Capabilities will expand over time, and your feedback helps shape what we build next.
Overview
Spatial transcriptomics lets you measure gene expression across a tissue section while preserving the physical location of each measurement. Instead of just knowing which genes are active, you can see where in the tissue they are active, mapping gene activity directly onto your tissue image.
In Pluto, Spatial Transcriptomics is a dedicated experiment type that lets you upload your processed data and explore it interactively using a built-in spatial browser, powered by Vitessce. No coding required.
Because Pluto uses Vitessce as its visualization engine, a compatible tissue image file is required alongside your gene expression data. The sections below explain exactly what you need and how to get set up.
Before you begin: What to prepare
Pluto expects your spatial data to be fully processed before upload. This means your bioinformatics team (or your pipeline) should have already completed:
Quality control (QC): low-quality spots and outlier cells removed
Normalization: counts normalized across spots
Clustering: spots grouped into clusters (e.g., Leiden, Louvain, or custom annotations)
Dimensionality reduction: embeddings such as UMAP or PCA computed
Spatial coordinates: X/Y tissue positions stored in the object
If your data has not yet been processed, work with your bioinformatics team before uploading to Pluto.
⚠ Single sample per experiment: Pluto currently supports one sample and one tissue image per Spatial Transcriptomics experiment. If you have multiple samples, create a separate experiment for each one.
Required files
You will need two files to set up a Spatial transcriptomics experiment in Pluto:
File | Format | What it contains |
Spatial object | .h5ad (AnnData) | Processed gene expression counts, spatial coordinates, cell metadata, cluster annotations, and embeddings |
Tissue image | OME-TIFF (.tiff / .tif) | High-resolution tissue image associated with your spatial object, required by the Vitessce viewer |
📌 Both files are required. Pluto will prompt you to first upload your .h5ad file, and will prompt you later upload the OME-TIFF separately during experiment set-up.
About the .h5ad file
The .h5ad format (AnnData) is a single file that bundles gene expression counts, spot-level metadata, spatial coordinates, and embeddings together. It is the standard output format from spatial transcriptomics analysis pipelines built on Scanpy or similar tools.
At minimum, your .h5ad file must contain:
Gene expression matrix: spots × genes count matrix (raw or normalized)
Spatial coordinates: stored in obsm['spatial'] or equivalent
Cell/spot metadata: at least one column that can serve as a Sample ID, and at least one clustering column
Embeddings: dimensionality reductions such as UMAP or PCA (stored in obsm); these will be available for selection during the mapping step
About the OME-TIFF file
Pluto uses Vitessce to render your tissue image alongside gene expression data. Vitessce requires the image to be in OME-TIFF format. OME-TIFF files store high-resolution microscopy images in a tiled, multi-resolution structure that allows smooth navigation at different zoom levels.
OME-TIFF files are typically produced by your imaging instrument or by tools like bfconvert, QuPath, or the 10x Genomics Space Ranger pipeline. If you are unsure whether your image is in OME-TIFF format, check with your imaging core or bioinformatics team.
Uploading your data: Step-by-step
Setting up a Spatial Transcriptomics experiment in Pluto takes three steps, reflected in the progress sidebar you will see on screen: Create your experiment, Add your assay data, and Map your data.
Step 1: Create your experiment
Log into your Pluto workspace.
Create a new experiment and select Spatial Transcriptomics as the experiment type, along with the appropriate organism.
Be sure to give your experiment a descriptive name, and select the Project you want the experiment it live under.
Step 2: Upload your data (.h5ad and OME-TIFF image)
In the experiment wizard, click Choose file to select you .h5ad file from your computer.
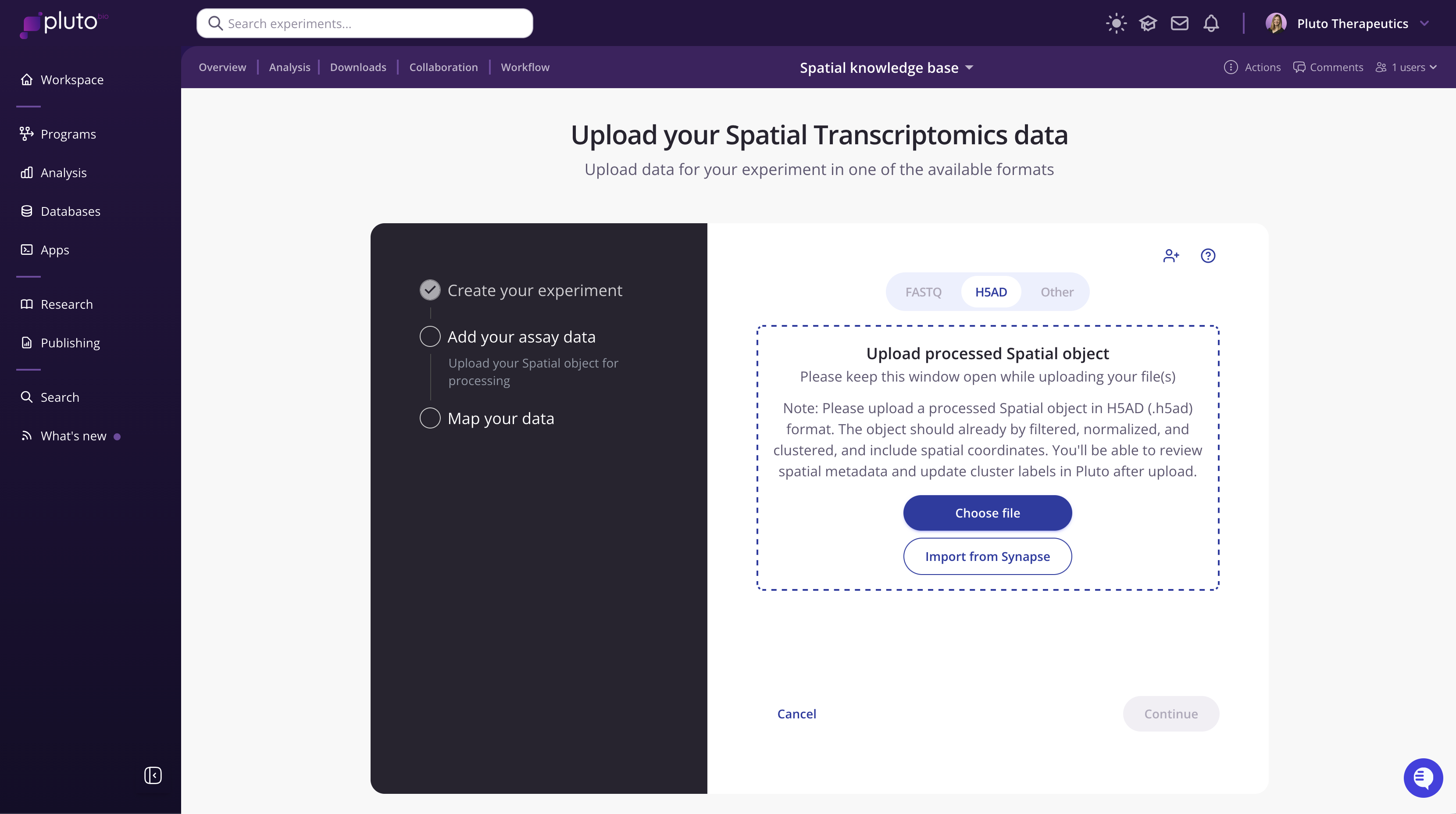
Once the upload is complete, Pluto will extract metadata and embedding information from your object automatically to use for mapping (Step 3).
Click Continue to proceed. You will be prompted to upload your OME-TIFF. Once the OME-TIFF has been uploaded, you will guided through mapping your data.
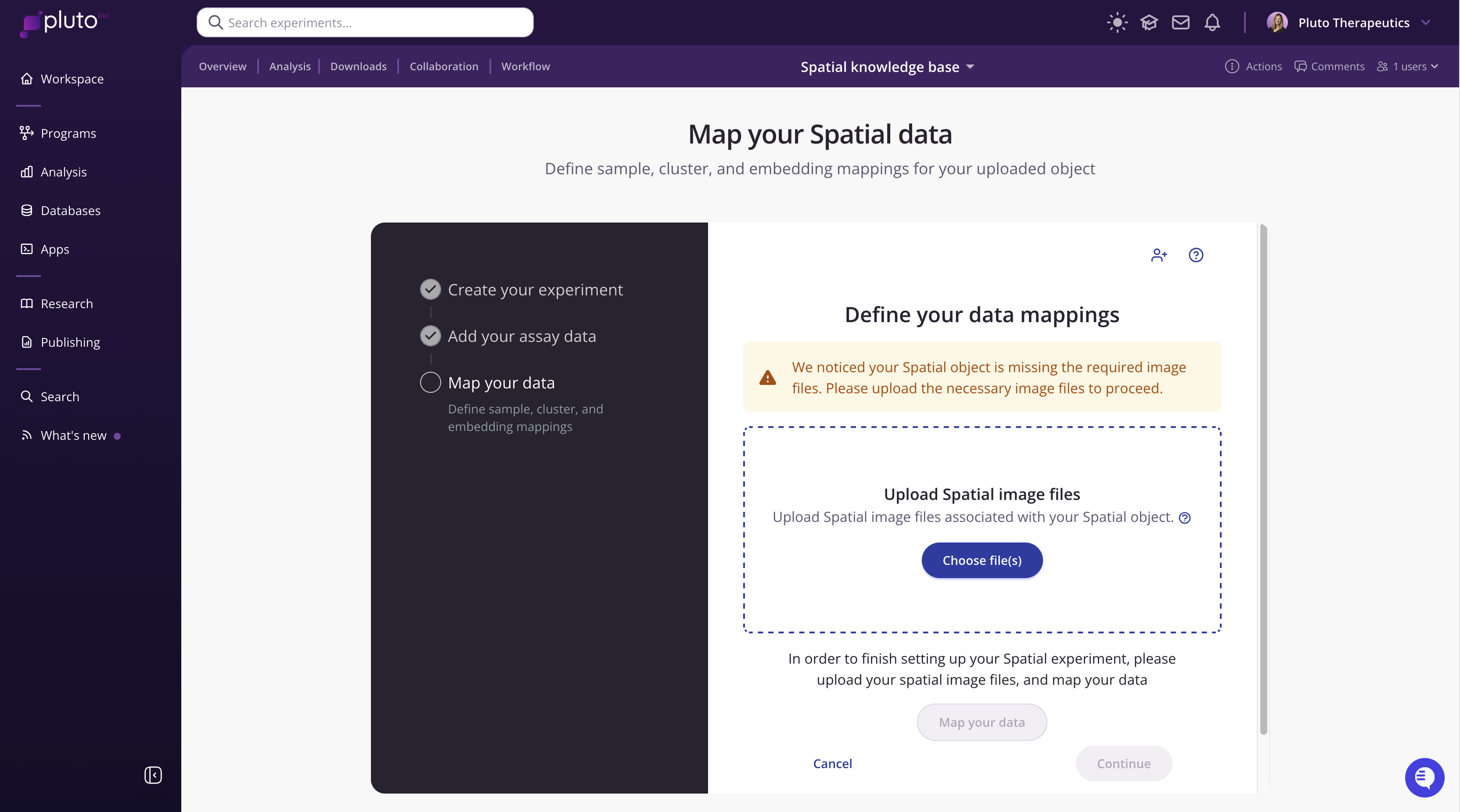
Step 3: Map your data
Once your assay data is uploaded, click Map your data to open the mapping dialog. This is a two-part process: first you will map your cell metadata, then configure your embeddings.
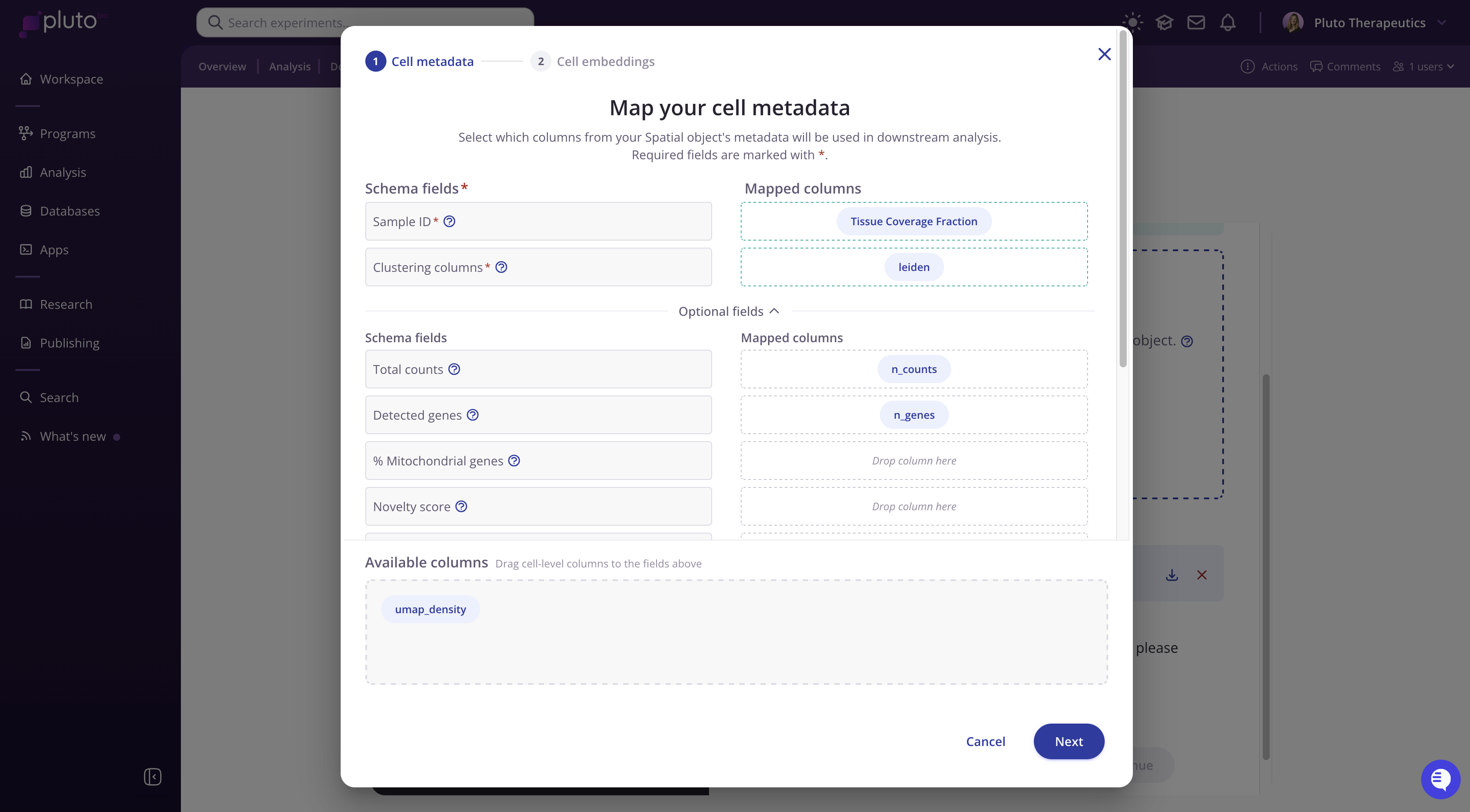
Map your cell metadata
Pluto will display the columns detected in your .h5ad metadata as well as our schema fields.
Note that required fields are marked with *:
Sample ID: The column in your metadata that identifies each sample (e.g., "sample_id", "library_id", etc.).
Clustering columns: One or more columns containing cluster or cell type annotations (e.g., "leiden", "cell_type").
Optional schema fields can additionally be mapped.
To map a field, simply drag the available column chips from the Available columns tray into the appropriate field boxes. Pluto may auto-match some columns, so be sure to review those before proceeding.
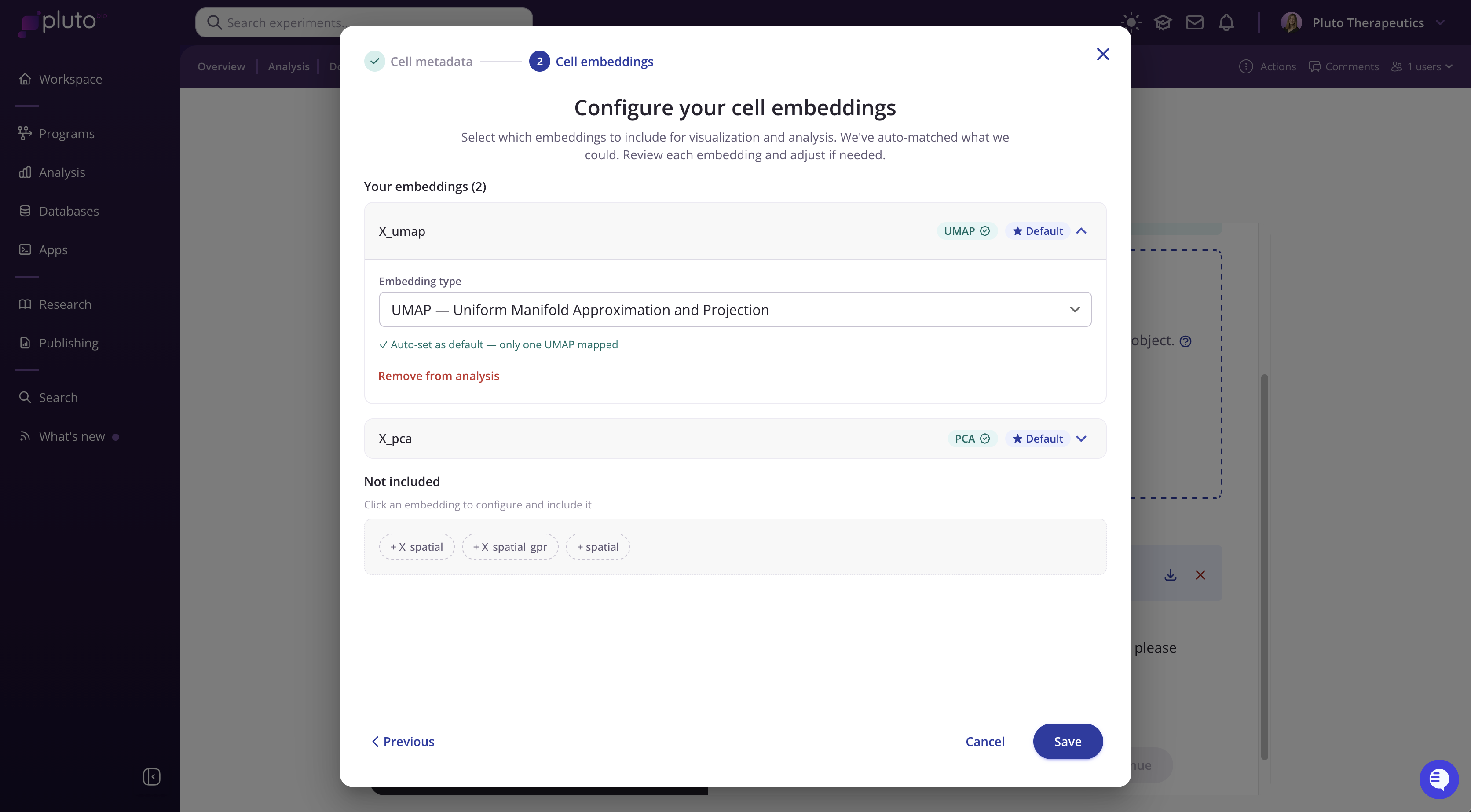
Configure your cell embeddings
Pluto will display the embeddings it detected in your .h5ad file. It will attempt to auto-identify common embedding types such as UMAP, tSNE, and PCA.
Included embeddings (shown at the top) will be available for visualization. You can review and change the embedding type label using the dropdown.
Not included (shown at the bottom) are embeddings Pluto found but did not auto-include. Click any of these to configure and add them if desired.
Spatial coordinate embeddings (e.g., X_spatial, spatial) may appear in the Not included section. These are used by the Vitessce viewer and do not need to be mapped as analysis embeddings.
Click Save to finalize your mappings and submit your data for processing and Pluto-integration.
Exploring your data
After saving your mappings, Pluto will run a pipeline to integrate your .h5ad object and OME-TIFF image into the platform. This may take a few minutes depending on the size of your dataset.
Once the pipeline is complete, you will be able to explore your data using our in-app Vitessce browser.